In our lab, we characterise bacteriophages to be used as antimicrobial therapeutics, known as Phage Therapy (read more). Two aspects that are directly relevant for the clinical use of bacteriophages are of particular interest to our group:
(1) Bacterial Phage-Resistance, i.e. the emergence of strains that cannot be infected by a phage, often due to the loss of a receptor, among various other mechanisms. Phage resistance is not the end of a therapeutic approach but can be advantageous, as such resistant strains often re-sensitise to antibiotics and show a decrease in virulence.
(2) Phage-Antibiotic-Interaction, often referred to as PAS, or Phage-Antibiotic-Synergy, as phages often show cooperative effects with antibiotics. This even occurs, when the bacterial pathogen is fully resistant to the antibiotic compounds, and is not fully understood.
In addition, we investigate fundamental aspects in biology such as the function of phage proteins, which can display antimicrobial, anti-biofilm or anti-capsule properties. We are also interested how phages function, how they interact with the host and how they assemble.
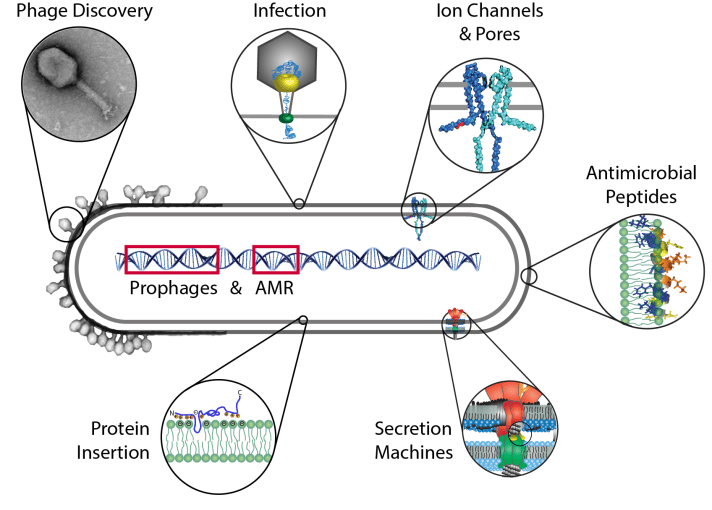
The major topic of our research is the membrane of cells. The membrane is the border that separates the outside from the inside of the cell. However, the cell envelope and lipid bilayer is much more than a sack that surrounds the cell. Molecules and also signals have to cross this barrier. Here it is also, where phages bind and interact, and facilitate the transport of their genome into the host, often called a DNA “injection”.

In our group we often study membrane proteins of microbes which are of a much simpler molecular architecture and thus easier to understand than their human counterparts. We investigate how such proteins insert, assemble and function in the membrane.
We also study proteins and peptides that are secreted and interact with the membrane of a cell that is being attacked by e. g. a pathogen. An example would be bacterial secretion systems that make use of this mechanism to assemble a structure much like a syringe, to inject molecules into the eukaryotic host. Another interest in our group is the investigation of how membrane-embedded molecular motors works.
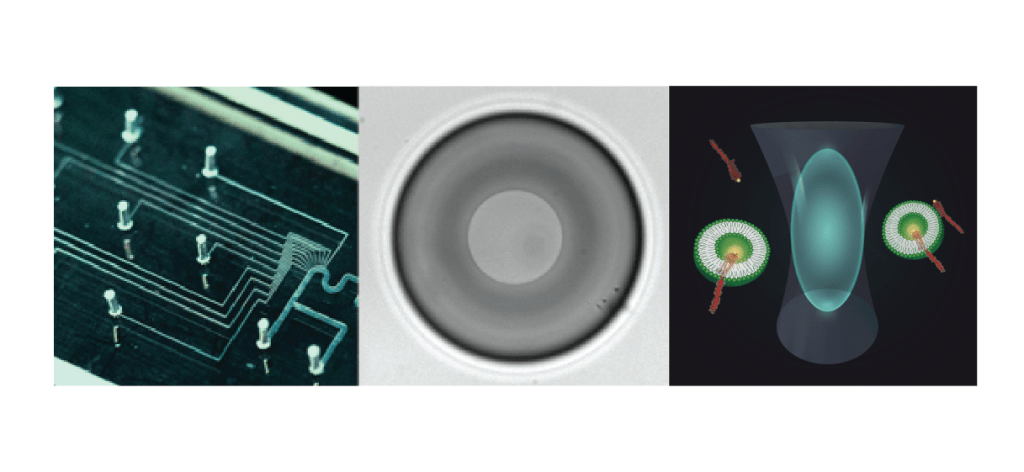
We make use of a variety of methods, such as microfluidics, artificial lipid bilayer techniques and single molecule fluorescence. Microfluidics (above, left), in particular the use of so-called water-in-oil droplets, together with powerful imaging and sorting techniques, allow fast screening for many purposes: Studying phage-bacteria interaction, bacterial evolution, compound screening, but also protein activity assays and so much more!
Many of our projects start with traditional methods that include microbiology, molecular biology and biochemistry techniques. To understand protein function or interaction in the membrane, we use single molecule methods such as single molecule fluorescence and single channel electrical recording. Both are mostly done in artificial bilayers systems. With the technique of the Droplet-Interface-Bilayers (DIBs – middle), the group has a fantastic tool to perform experiments using electrical and optical measurements simultaneously, including single molecule TIRF, smFRET, and FCS (above, right).
In addition to membrane biochemistry, we also work with bacteriophages, or viruses of microbes. We investigate two aspects: (1) We aim to understand virus assembly and DNA-packing of filamentous phages which are important for the phage display technique, and (2) we explore the aspects of phage discovery and therapy. Due to over-treatment and misuse, many pathogens have become resistant to chemical antibiotics, therefore phages are promising alternatives to inactivate bacteria that cause infections.
Due to the coronavirus pandemic and many of our lab members not being able to work at the bench, we started several bioinformatics projects together with our collaborators. We are very happy to be able to work together with Dr. Hua from the Shaw hospital on genome analysis of prophages and AMR. Several exciting projects have also been started with bioinformatician, Dr. Melanie Stefan from the University of Edinburgh (Centre for Integrative Physiology) where we model phage infection and dynamics.